How to Visualize PCA Results in R, An unsupervised machine learning technique called principal components analysis (PCA) looks for primary components that account for a significant amount of the variation in a dataset.
A biplot, which plots every observation in a dataset on a plane defined by the first two main components, can be used to see the PCA findings for a particular dataset.
To make a biplot in R, we can utilize the fundamental syntax shown below:
How to Visualize PCA Results in R
We’ll use the USArrests dataset from the built-in R environment for this example.
See the USArrests dataset’s first six rows.
head(USArrests)
Murder Assault UrbanPop Rape Alabama 13.2 236 58 21.2 Alaska 10.0 263 48 44.5 Arizona 8.1 294 80 31.0 Arkansas 8.8 190 50 19.5 California 9.0 276 91 40.6 Colorado 7.9 204 78 38.7
To perform PCA and display the results in a biplot, we may use the code below:
How Do Machine Learning Chatbots Work – Data Science Tutorials
proceed to PCA
results <- princomp(USArrests)
PCA findings can be shown in a biplot.
biplot(results)

The first principle component is displayed on the x-axis, the second principal component is displayed on the y-axis, and the individual dataset observations are displayed inside the plot together with the four red-hued variables.
Be aware that the biplot function has a number of arguments we can use to change the plot’s appearance.
For instance, we can change the plot’s colors, font size, axis boundaries, plot title, axis titles, and size of the arrows by using the following code:
Let’s generate a custom-looking biplot
biplot(results,
col=c('blue', 'red'),
cex=c(1, 1.3),
xlim=c(-.4, .4),
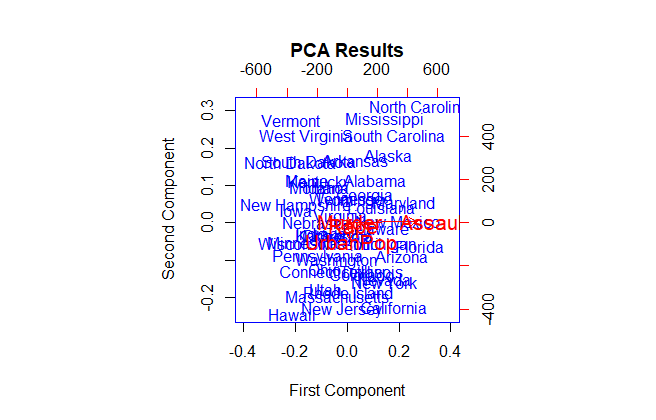
main='PCA Results',
xlab='First Component',
ylab='Second Component',
expand=1.2)
It’s a little bit simpler to interpret this biplot than the last one.